Using this module, it is possible to define any array of named characters, binary or continuous, with fixed or undefined length. The size of a character experiment in BIONUMERICS can range from one single character (e.g. a morphological feature) to microarray experiments of many thousands of gene expression values. There is no limit to the size of the character array and the software is optimized to analyze large data sets.
Examples of character data include biochemical and morphological features, metabolic assimilation or enzyme activity test panels such as API®, Biolog®, Vitek®, antibiotics resistance profiles, fatty acid profiles, microarray differential expression values, etc.
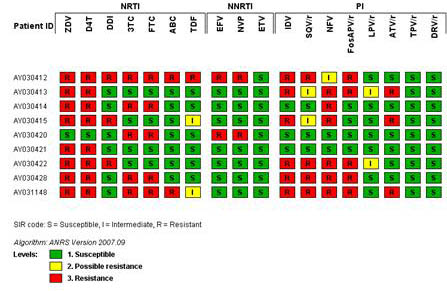
Character sets are also used by plugin tools to store the result of processed data from other sources, for example, VNTR copy numbers in Multi-Locus VNTR Analysis (MLVA), allele numbers in Multi-Locus Sequence Typing (MLST), zygosity in AFLP band scoring, resistance profiles in antibiotic resistance profiling in bacteria and HIV drug resistance analysis, etc.
Character import from a text file or an Excel spreadsheet
BIONUMERICS has very flexible and powerful import tools for the import of character data from a text file (e.g. in tab-delimited or CSV format), Microsoft Excel spreadsheet and any ODBC-compatible database such as Microsoft Access.
Customized display of test panels
Character profiles can be displayed in a panel with user-defined representations and color scales or in a list with values. The character display tools in BIONUMERICS are very flexible, resulting in a truthful image of all test results, be it from a commercially available test panel (e.g. Biolog plates, bioMérieux' API strips), DNA or microarray, microplate assay (e.g. ELISA tests in 96 or 384 well plate format) or any standard biochemical test.
Distance and similarity matrices
An externally generated distance matrix or similarity matrix can be imported and linked to database entries in a BIONUMERICS database. This is used in conjunction with other information to obtain classifications and identifications. The distance or similarity values are either measured directly by the technique (a typical example being DNA-DNA hybridization values in bacterial taxonomy), or generated by other software.
